New updates and news from the team at Pluto. Here's the latest new Pluto feature run-down✨
Introducing comments & notes
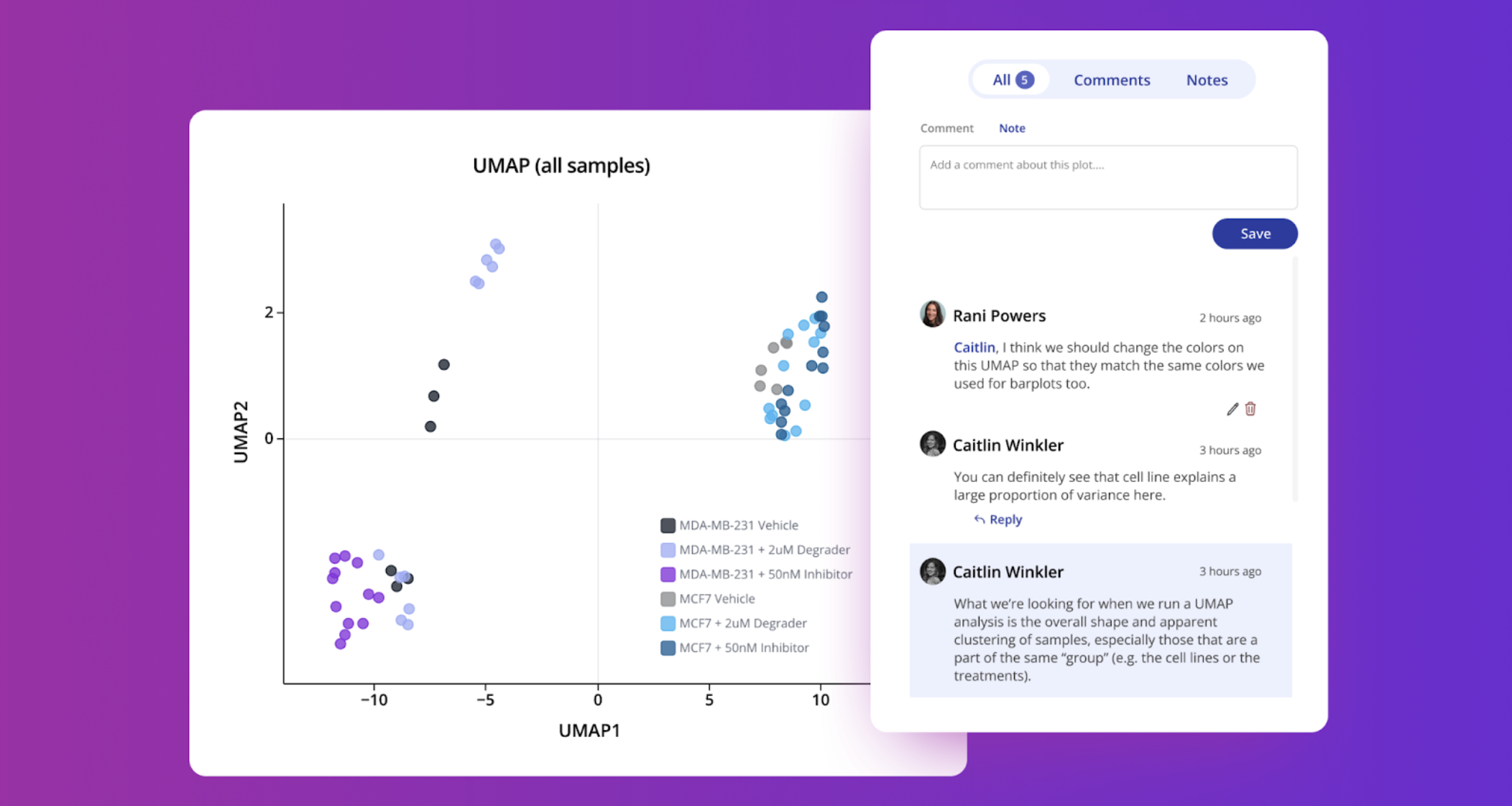
Collaborate with your team in real time - leave comments on experiments or individual plots to capture feedback, and use research notes to record your team's important thought processes and biological interpretation.
Covariates in differential analyses
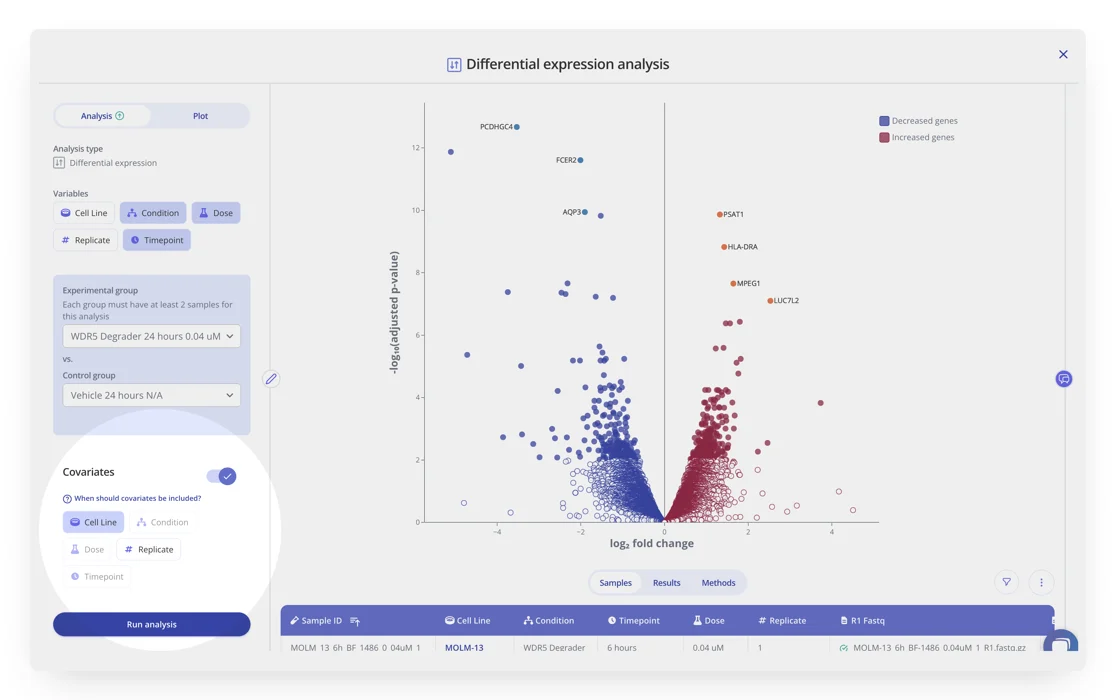
When comparing samples, it's important to take into account confounding variables, such as batch, age, sex, and other potential covariates. Now, when running differential expression analyses in Pluto, you have the option to include covariate(s) in the model.

Researchers from the Wyss Institute at Harvard University used Pluto to analyze RNA-seq data to investigate acute radiation induced lung injury. We sat down with the lead author to discuss the impact of this exciting work, and how Pluto accelerated the analysis and visualization of their complex, high-throughput data sets.
In case you missed it...
In the past month, we also launched:
- Export to Microsoft Powerpoint and Word
- Gene set enrichment analysis for epigenetics experiments (ChIP-seq, CUT&RUN)
- Updates to the Pluto R package and Python SDK- contact your Pluto rep for a personalized training!
Haven't started using Pluto's collaborative computational biology platform yet?
Whether you're analyzing data in real-time with colleagues, or securely collaborating with external partners across the globe, Pluto empowers you to gain insight from data, create presentation-ready plots, and accelerate your team's R&D workflow.
Stay tuned!
We’re launching even more highly requested features every month.
Thanks for reading! Say hello any time at hello@pluto.bio - we’d love to hear your thoughts on the features we’re highlighting this month.

